Researchers identified gene regulatory subcircuits controlling neocortical excitatory neuron (ExN) specification, with ZBTB18-CRE interactions playing central roles in mammalian-specific adaptations. Using transgenic mice expressing GFP in upper layer intratelencephalic (IT) neurons or deep layer extratelencephalic (ET) neurons, scientists enriched these populations from neonatal cortex and performed RNA sequencing and chromatin immunoprecipitation assays. Analysis revealed transcription factor binding sites enriched in IT-biased and ET-biased regulatory regions. The study identified 69 transcription factors with differential enrichment between neuron subtypes, establishing molecular mechanisms underlying neocortical ExN identity, connectivity, and laminar organization that distinguish mammals from other vertebrates.
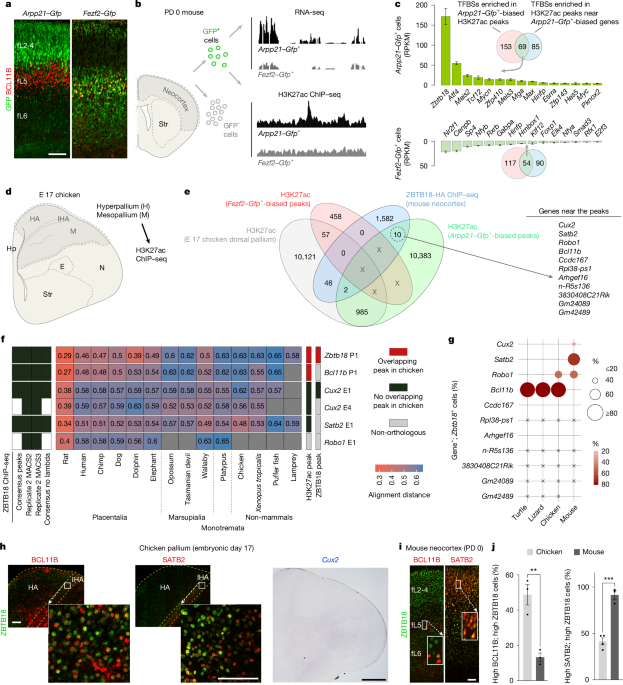
"To characterize CREs and TFs for neocortical ExNs, we used Arpp21-Gfp or Fezf2-Gfp transgenic mice and enriched GFP-expressing neocortical upper layer (L2-4) intratelencephalic (IT) neurons or deep layer (L5-6) predominantly extratelencephalic (ET) neurons, respectively, from neonatal mice (postnatal day (PD) 0), an age at which neocortical ExN identity and connectivity are established."
"Here we uncovered gene regulatory subcircuits, particularly involving ZBTB18-CRE interactions that govern key features of neocortical ExNs that have undergone modifications in the mammalian lineage."
"TFBS for 69 TFs enriched among both IT-biased H3K27ac peaks and H3K27ac peaks near IT-biased genes compared with ET-biased peaks and genes, respectively, with significant TF motifs having a Benjamini-Hochberg-corrected P value < 0.05 and odds ratio > 1."
#neocortical-excitatory-neurons #transcription-factors #gene-regulatory-circuits #mammalian-evolution #chromatin-regulation
Read at Nature
Unable to calculate read time
Collection
[
|
...
]