"Igh and Igk V(D)J recombination is orchestrated by the RAG1-RAG2 heterotetramer bound to a recombination centre formed around intronic enhancers and J segments of each locus. RAG cleaves robustly only paired gene segments flanked by bona fide RSSs with, respectively, 12-bp and 23-bp spacers, which must be properly aligned in parallel in the active sites of the two RAG1 proteins in the RAG complex."
"Cohesin-mediated chromatin loop extrusion contributes to pairing widely separated Igh and Igk RSSs for V(D)J recombination. For Igh, impeded downstream loop extrusion at the RC leads to continued extrusion of upstream chromatin through the impeded cohesin ring, allowing J H-23RSS-bound RAG to linearly scan the upstream 2.5 megabases of chromatin for D-12RSSs and, ultimately, V H-23RSSs."
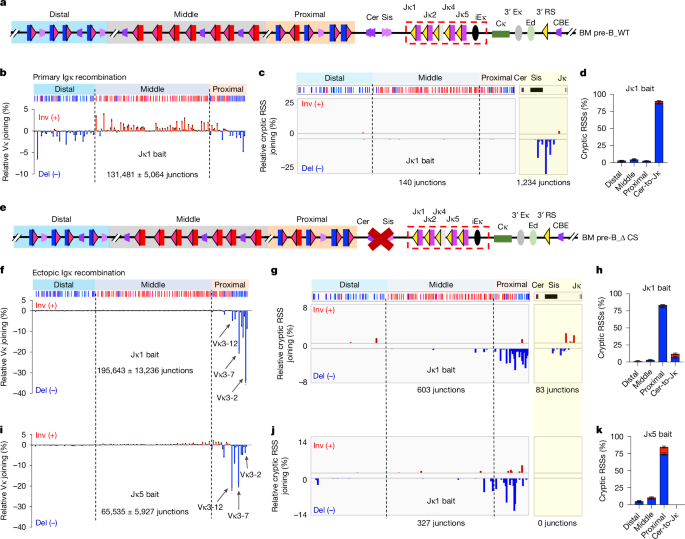
"The mouse Igk has 4 Jκs with upstream-oriented 23RSSs and more than 100 Vκs with 12RSSs, which lie in three clusters of mostly downstream oriented distal Vκs, mostly upstream oriented middle Vκs and both downstream and upstream-oriented proximal Vκs. Vκs across the 3.1 Mb locus are robustly used by the Vκ-to-Jκ1 joining process that generates initial VκJκ1 repertoires."
V(D)J recombination is facilitated by the RAG1-RAG2 complex, which binds to recombination centers formed around intronic enhancers and J segments. Recombination signal sequences (RSSs) flank Vs, Ds, and Js, enabling RAG endonuclease activity. RAG cleaves paired gene segments with specific spacers, requiring proper alignment. Cohesin-mediated chromatin loop extrusion aids in pairing distant RSSs. The mechanism allows RAG to scan chromatin for D-12RSSs and V-23RSSs. Mouse Igk has multiple Jκs and Vκs, with specific orientations affecting the joining process.
Read at Nature
Unable to calculate read time
Collection
[
|
...
]