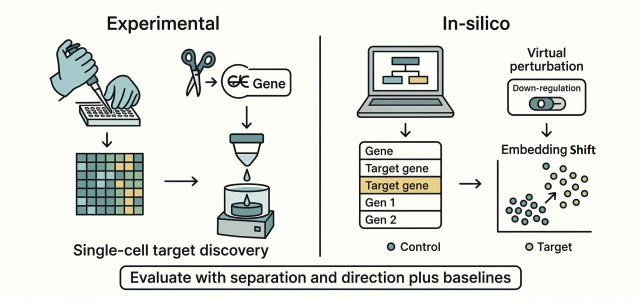
"Single-cell foundation models like Geneformer and scGPT are pretrained on large single-cell atlases, enabling them to learn embeddings applicable to various downstream tasks with minimal additional modeling effort."
"The gap between the technical possibility of in-silico perturbation and its biological trustworthiness is significant, as many evaluations do not align with critical discovery questions."
"Recent studies, ISP1 and ISP2, advocate for a benchmark-first approach, emphasizing the validation of state transitions before placing trust in simulated perturbations."
In-silico perturbation (ISP) simulates cellular state changes due to interventions like genetic or chemical manipulation. Single-cell foundation models (scFMs) such as Geneformer and scGPT, pretrained on extensive single-cell atlases, facilitate this process. However, a gap exists between technical capabilities and biological reliability. Evaluations often focus on transcriptome reconstruction, which may not align with discovery objectives. Recent studies suggest validating state transitions before trusting simulated perturbations, emphasizing the need for accurate assessments of how perturbations influence cellular states.
#in-silico-perturbation #single-cell-foundation-models #geneformer #scgpt #biological-trustworthiness
Read at Medium
Unable to calculate read time
Collection
[
|
...
]